NIBIB-funded researchers are building macroscopic medical images of cells and tissues at speeds 7,000 times faster than current methods allow, using a deep learning approach. The production of high-quality, real-time macroscopic molecular images could allow medical experts to make better and more informed clinical diagnoses. The new technique, developed by a team at Rensselaer Polytechnic Institute, can produce better images of living organisms in real time.
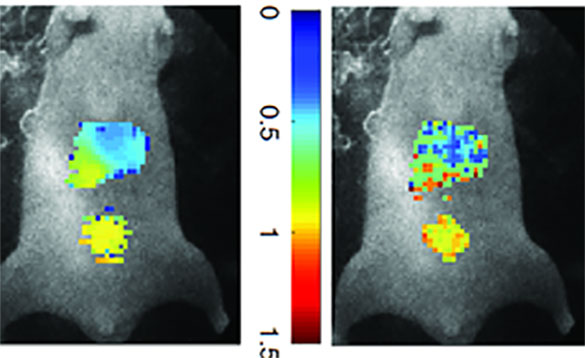
Molecular imaging allows medical experts to see a detailed picture of what is happening inside a person’s cells and tissues by exploiting various traits of molecules associated with a disease, also known as biomarkers. In optical imaging, scientists place fluorescent labels on the biomarkers.
They can simultaneously monitor multiple biomarkers by varying the fluorescent colors attached to each, making them easy to differentiate from each other. With this technique, they can measure the time that a molecule remains glowing or in an excited state, which is defined as “lifetime.” Measuring the lifetime of a molecule can be useful for monitoring drug delivery or the environment immediately surrounding the molecule (microenvironment).
The time it takes to process data “is one of the main barriers in measuring the lifetime of multiple fluorescent biomarkers, especially for clinical applications,” said Behrouz Shabestari, Ph.D., director of the NIBIB program in Artificial Intelligence, Machine Learning and Deep Learning. “Advances in artificial intelligence in biomedical imaging, especially deep learning, are beginning to significantly improve image quality and speed up processing times.”
Deep learning is a subset of machine learning that is a set of artificial neural networks, often compared to the human brain. A network consists of many nodes, similar to neurons in the brain. An assigned weight amplifies or reduces a node’s input, depending on the assigned task.
Pingkun Yan, Ph.D., co-director of the Rensselaer Biomedical Imaging Center, explained: “Nodes are the sites where the computation takes place; they are like filters in convolutional neural networks.” He described convolutional neural networks as many layers of nodes that are stacked on top of each other, and data is fed through the network one layer at a time; But, like neurons in the brain, not all nodes activate for each task. The network must be trained with a large amount of data to be successful in processing information.
Deep learning has been applied to biomedical imaging to improve the quality of an existing image and improve clinical diagnosis. “But our team’s goal is unique; we are using deep learning to perform the physics behind the original construction of the image,” explained Xavier Intes, Ph.D., co-director of the Rensselaer Biomedical Imaging Center and corresponding author of the paper published in the journal Nature. Light: science and applications.
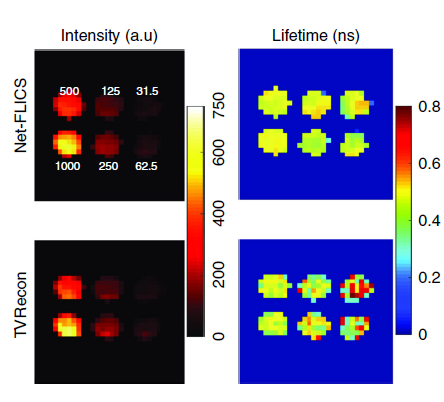
Building on a previous approach developed by the same team, the new technique called Net-FLICS (fluorescence lifetime imaging with compressed sensing) uses deep learning to construct an image from a small number of point measurements.
Intes and Yan’s team created a completely new deep learning network to create molecular images. Often, a network that is exceptional at identifying images in a Google search can be slightly tailored for similar applications such as facial recognition; This process is called transfer learning. “There was no network for our specific application, so we had to take pieces from other networks to build what we needed,” Yan said.
The researchers trained the network using simulations and computer models to generate a large amount of data. Using models gave the team a huge advantage: they had access to an infinite amount of data, which is a common obstacle in training a network.
However, there are challenges when working with a new network. “It’s hard to understand what’s happening inside a new network like ours; it’s like a black box. As we continue to learn more, it will become clearer why the network makes certain decisions and we can improve them,” Intes explained.
The significant advantages of Net-FLICS enable near real-time image acquisition, opening new doors for researchers and clinicians due to the improved speed and quality of images. Future applications of the Net-FLICS technique will help identify which individual cells show acceptance of drug treatment and which do not. Real-time molecular imaging could also help surgeons while performing complicated procedures such as removing a tumor or working on delicate nerves.
This work was supported by National Institutes of Health grants R01 EB19443 and R01 CA207725.
Yao R, Ochoa M, Yan P, Intes X. Net-FLICS: Rapid Quantitative Wide-Field and Lifetime Fluorescence Imaging with Compressed Sensing: A Deep Learning Approach. Light science app. 2019; 8:26. Published 2019. doi:10.1038/s41377-019-0138-x